Alex completed his PhD in biology at UPenn, working in Dustin Brisson’s lab where he studied the population genetics of Trypanosoma cruzi in Peru. He was a postdoc with us from 2018-2020, during which time he instrumental in our studies of the microbiome in large animal models. He was the first to show that natural parasite infections are a dominant factor in shaping microbiome structure in humans and animals (preprint here), and his most recent work used pigs to understand the impact of pregnancy and pregnancy history (parity) on the microbiome and offspring colonization (see here). In addition, Alex played a key role in supporting and developing our transcriptomics course, and was one of the lead authors on this perspectives piece that lays out our approach for teaching bioinformatics to bench biologists and how our curriculum was adapted to improve virtual content during the COVID19 pandemic.
Elise was our lab manager and sequencing guru from 2019-2021, and she played a vital role in helping the lab survive and continue to operate during the worst of the COVID19 pandemic. She left the lab to pursue a PhD in Biology at the University of Syracuse.
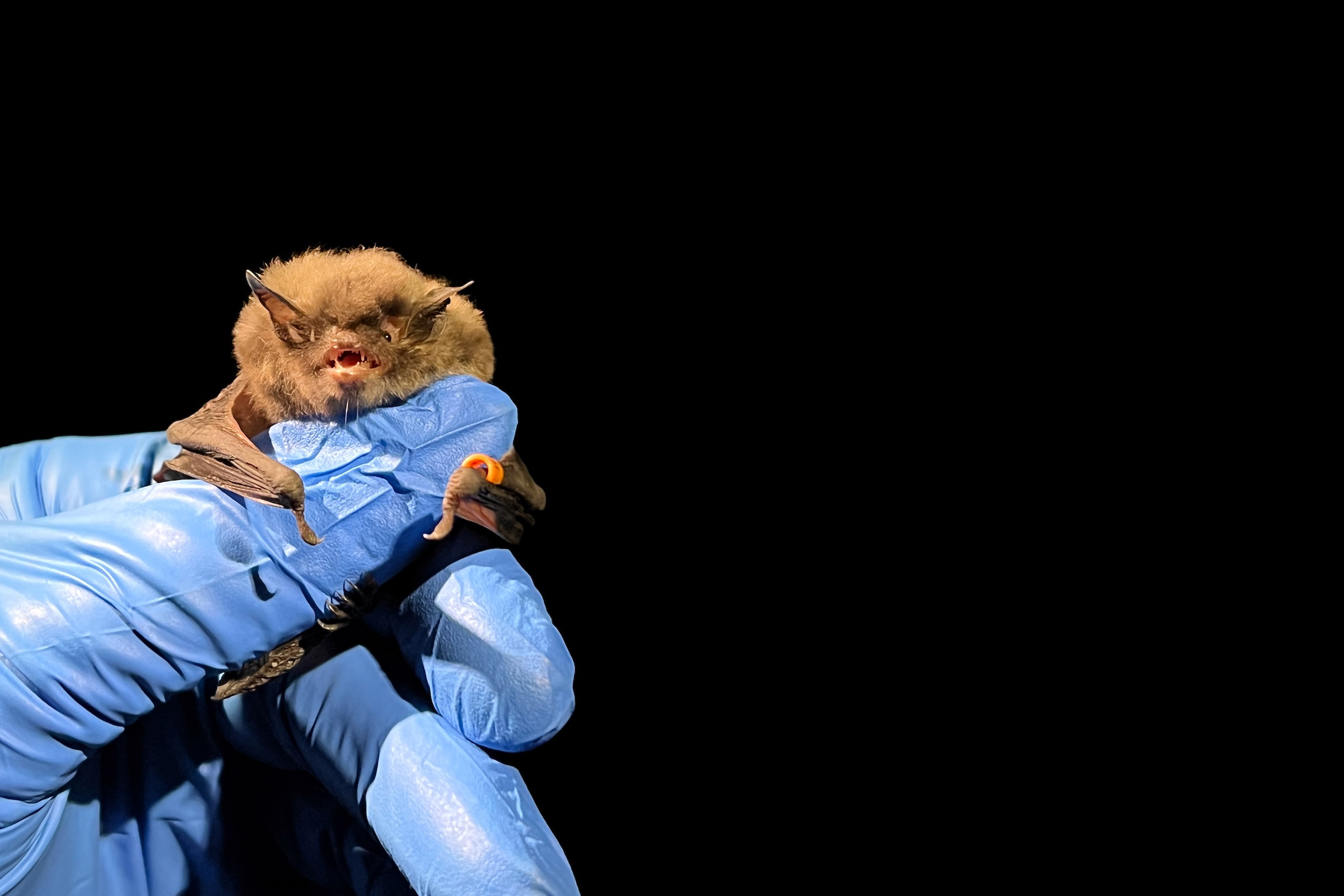
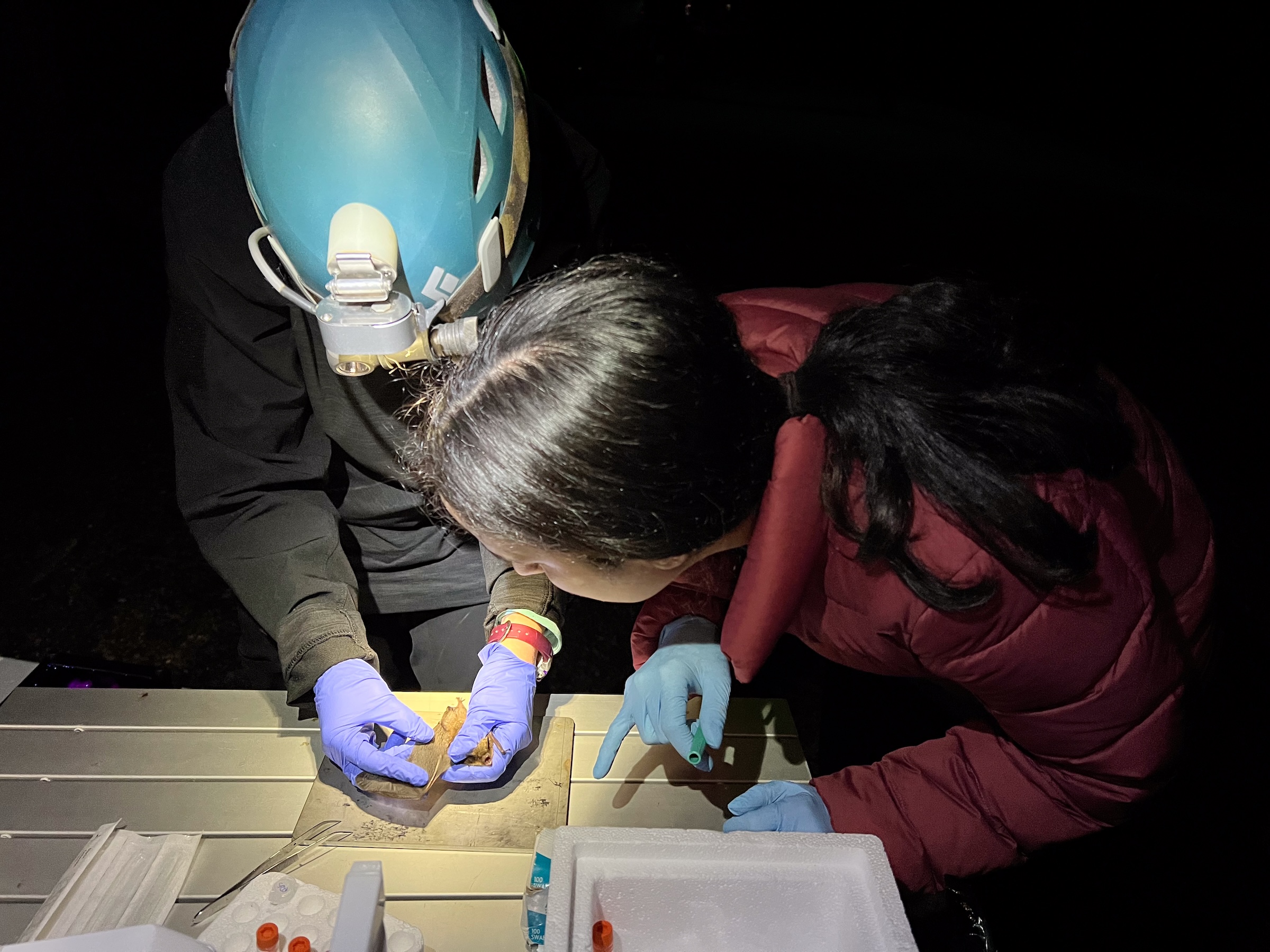
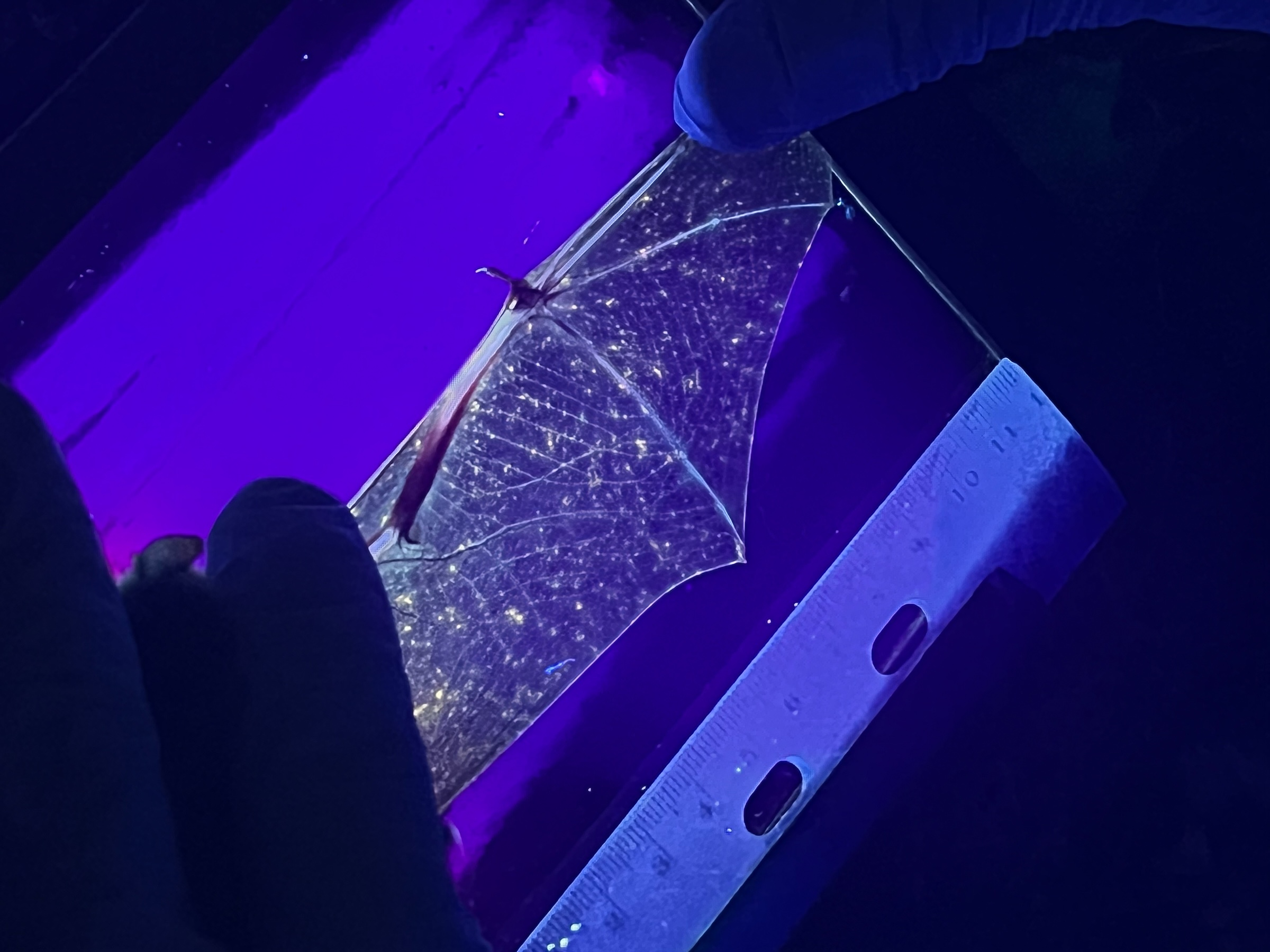
Clara came to us straight out of her undergraduate studies at Smith College, where she worked in the laboratory of Dr. Laura Katz. After learning the ropes, Clara became an incredibly effective manager of all high-throughput sequencing projects provided as a service by the lab. Clara also spent time doing field work and made critical contributions to our metagenomics work in bats affected by White Nose Syndrome. After two years as lab manager, Clara left the lab to start her Graduate program in the CAMB/MVP program at UPenn. Lucky for us, we still get to see Clara every day – she ended up joining the lab of Louise Moncla, our lab neighbors and with whom we share a weekly lab meeting. We’re so proud of her and can’t wait to see all that she accomplishes in the Moncla lab!
For nearly 20 years I have studied host-microbial interactions, with a primary focus on leveraging genomic and bioinformatic approaches to elucidate mechanisms that regulate inflammation and host defense during infection with zoonotic microbes. During my PhD in Judy Appleton’s lab at Cornell University’s Baker Institute for Animal Health, I studied the immune response during chronic infection with the parasitic helminth, Trichinella spiralis. This work led to the identification of the cytokines IL10 and TGF-β, and eosinophils, as critical regulators of inflammation and immunity to chronic helminth infection. As as postdoc in David Roos’ lab at UPenn, my work shifted from helminth to protozoan parasites, and I began to employ genome-wide transcriptional profiling and genetic screens to identify novel players involved signaling pathways that regulate both immunity and pathogenic inflammation. The results of these studies helped shed light on important immune effector mechanisms, ranging from IL27 signaling in Tregs, to identifying novel enhancers of STAT1 signaling, to reactive oxygen species production by infected monocytes, to TLR3-dependent type I interferon production. In January of 2013, I began a faculty position in the School of Veterinary Medicine at UPenn, were I Co-founded and continue to Co-direct the Center for Host-Microbial Interactions (CHMI). My primary research mission in this role is to develop interdisciplinary ‘One-health’ projects related to microbiology and infectious diseases. In parallel with my research program, I work to engage trainees at the University level in bioinformatics training for managing and analyzing genomic datasets.
Amanda is a PhD student in the Immunology Graduate Group (IGG) and spent her 1st rotation with us. The goal of Amanda’s project was to interrogate the host transcriptional response during infection-induced dysbiosis, and to identify the microbes or microbial factors associated with inflammation leading to sepsis. Amanda used an infection with Toxoplasma gondii to induce severe dysbiosis fand collected gut tissue, stool and created a microbial isolate collection from mice at 0, 2, 5, and 8 days post-infection. Amanda then worked to isolate RNA from these tissues and prepare sequence-ready libraries. The goal of this project is to try to use RNAseq to simultaneous profile host and microbial gene expression during dysbiosis. This data set will provide insight into microbial community structure AND function during dysbiosis, as well as profiling the host response…all from the same sample.
Jake is a PhD student in the Genomics and Computational Biology (GCB) graduate group and spent his 2nd rotation with us. He employed ‘hybrid’ (short + long read) genome assembly methods to generate one of the first complete genomes for Clostridium hiranonis, a bile-acid producing member of the gut microbiome that is implicated in maintaining gut health through the production of secondary bile acids. Jake then annotated this genome and used comparative genomic approaches to understand population genetics of C. hiranonis across animals and humans. Since there is only one other complete genome available for this organism, Jake turned to mining our shotgun metagenomic data from animal stool samples and identified ~20 samples for which the metagenomic reads provide at least 100x coverage of the C. hiranonis genome. He used contigs from the metagenome assembled genomes (MAGs) in his comparative analysis.
Shuai was a postdoc in the lab from June 2017 to October 2019, during which time he focused on using animal models to identify mechanisms that regulate dysbiosis. Using an infection model in mice, he showed that strong immune responses to infection can also promote pathogenic changes in gut microbiome composition. His first paper describing this work, showed that nitric oxide produced by macrophages during infection serves as a substrate for E. coli expansion in the small intestine, ultimately leading to barrier breakdown and subsequent sepsis. His second project in the lab used a canine model of diet-responsive inflammatory bowel disease (IBD) to dissect relationship between the microbiome and remission from IBD. His paper on this topic showed that diet therapy induced a structural change in the microbiome, marked by expansion of the bile acid producer, Clostridium hiranonis, and a comensurate decrease in levels of potential pathobionts such as E. coli and C. perfringens. He then showed that C. hiranonis was sufficient to ameliorate inflammation and E. coli expansion in a mouse model of GI disease. These findings set the stage for future work in the lab that will examine bile acid producers in modulating inflammation.
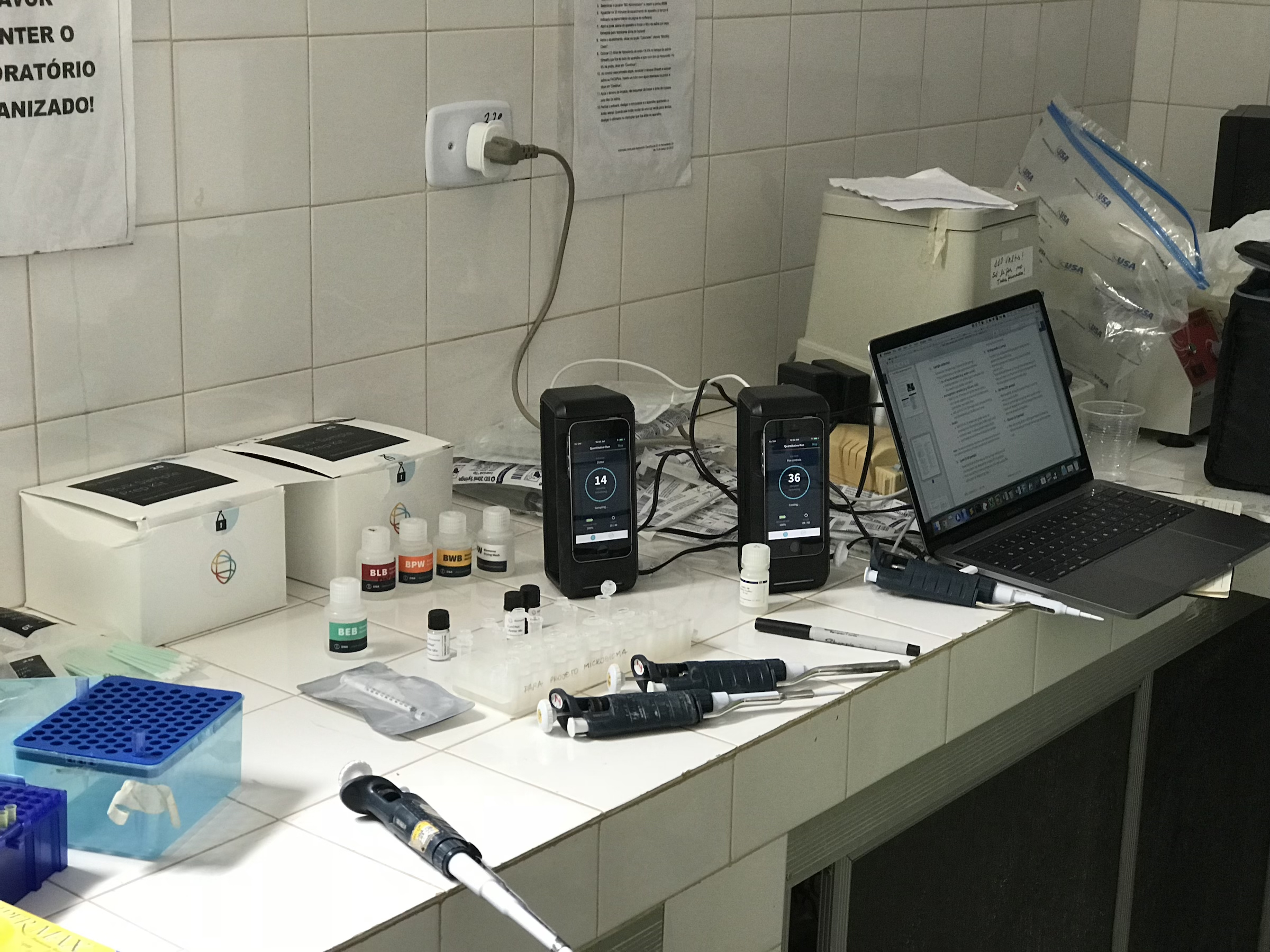
Matt (PennVet class of ‘22) spent the summer of 2019 in the lab as a NIH-BI research fellow. He continued work started by a previous student to optimize portable assays for detecting bacterial pathogens.
Megan joined the lab in October of 2017 and was a keystone of the lab until she left to start a master’s in biomedical sciences at PCOM in August of 2019. Megan handled all day-to-day operations in the lab, but her main contribution to the success of the our group involved managing all aspects of our high-throughput sequencing projects. This included everything from experiment planning; to executing technically challenging wet-lab experiments that transform raw biological samples into DNA suitable for sequencing; to operating sophisticated instrumentation used for sequencing; to troubleshooting failed experiments. During her time in the lab, she single-handedly sequenced over 1200 samples spanning over 100 projects. Megan has gone on to accomplish great things after leaving the laboratory. After completing her master’s degree, Megan got her Medical Degree at the Philadelphia College of Osteopathic Medicine (PCOM) and is now beginning her residency in orthopedic surgery. We never had any doubt that she was a superstar in the making!
Ayah was an undergraduate who carried out her honors thesis research in the lab until May of 2019. She was the first person in the lab to start doing E. coli competition experiments in mice infected with Toxoplasma gondii. Her data laid a rock solid foundation for work that ultimately led to this paper. After graduating with honors, Ayah worked in biotech for several years, and in 2022 she started her MD/MPH training at the University of Southern California’s Keck School of Medicine.
Pagination