Ann completed her Ph.D. and postdoc in the laboratory of Danielle Bassett at UPenn, where she used network theory to address questions in cognitive neuroscience. Ann’s role on the team is to advance data analysis and visualization methods for our microbiomeDB project, and she works closely with the larger VEuPathDB team that is our partner in this effort.
Qianxuan completed his first of three rotations in the lab and is a member of the Microbiology, Virology and Parasitology (MVP) Graduate group. Qianxuan’s project focused on a bile acid-prodcing commensal, Clostridium hiranonis, which we initially identified as an important factor in modulating GI disease. Due to COVID, Qianxuan has worked virtually to annotate a recently completed genome of C. hiranonis, while exploring published and unpublished shotgun metagenomic datasets to better understand this organism.
Seble spent the Summer of 2020 ‘in the lab’ (virutally, due to COVID) doing a rotation. She is a member of the Immunology Graduate Group (IGG), and her project focused on using bioinformatic tools to analyze single cell RNA sequencing data (scRNA-seq) from 3-dimensional cultures of intestinal epithelial cells in order to understand how these cells might be better poised to response to infection or inflammation.
Lydia spent the Spring of 2020 doing a virutal rotation in the lab during the earliest (and scariest!) phase of the COVID19 pandemic. She is a member of the Microbiology, Virology and Parasitology (MVP) Graduate group. Her rotation was mostly devoted to learning the computationoal methods and principles of RNA-seq.
Cleo was enrolled in the Masters of Biotechnology program at UPenn. As part of the molecular biology track within this program, Cleo did her independent study in the lab working with some of the bacterial isolates from our canine IBD study to try to set-up in vitro systems to study microbe-microbe interactions between bile acid producers and pathobionts. Unfortunately, this work was cut short by the COVID19 pandemic, but Cleo still finished her independent study virtually.
Alyssa spent her Junior and Senior years as an undergraduate researcher working in the lab. During this time, Alyssa simultaneous completed her Master’s degree in Biology, working to develop 3-Dimensional tissue culture models for studying how secondary bile acids impact immunity and barrier function. After graduating with her Bachelor’s and Master’s degrees in 2022 from UPenn, Alyssa started her PhD studies in Microbiology and Immunology at Columbia University.
Eman spent time in the lab when she was a senior at the University of the Sciences Philadelphia (USP), and spent the Spring semester of 2020 working in the lab on an independent study project. Eman was new to bench research and spent her time getting acquainted with fundamental techniques in molecular microbiology. Unfortunately, this work was cut short by the COVID19 pandemic, and Eman was accepted into a Postbac program at WashU.
Wojtek joined the lab in November of 2019 as our first bona fide software developer. He came to us from the Sanger Institute, where he served as the primary developer of Wormbase paraSite. His role on our team is to advance our microbiomeDB project, and he works closely with the larger VEuPathDB team that is our (essential!) partner in this effort. Wojtek moved on to other software development projects with this larger team, but during his time with us he loaded large enteric disease datasets into microbiomeDB, built private user workspaces that allow users to directly upload and analyze their own data, developed an automated workflow that enables handling of shotgun metagenomic data, and built a new software tool (CORRAL) that allows investigators to identify microbial eukaryotes in complex metagenomic datasets. Wojtek now works as an independent compsci and compbio consultant.
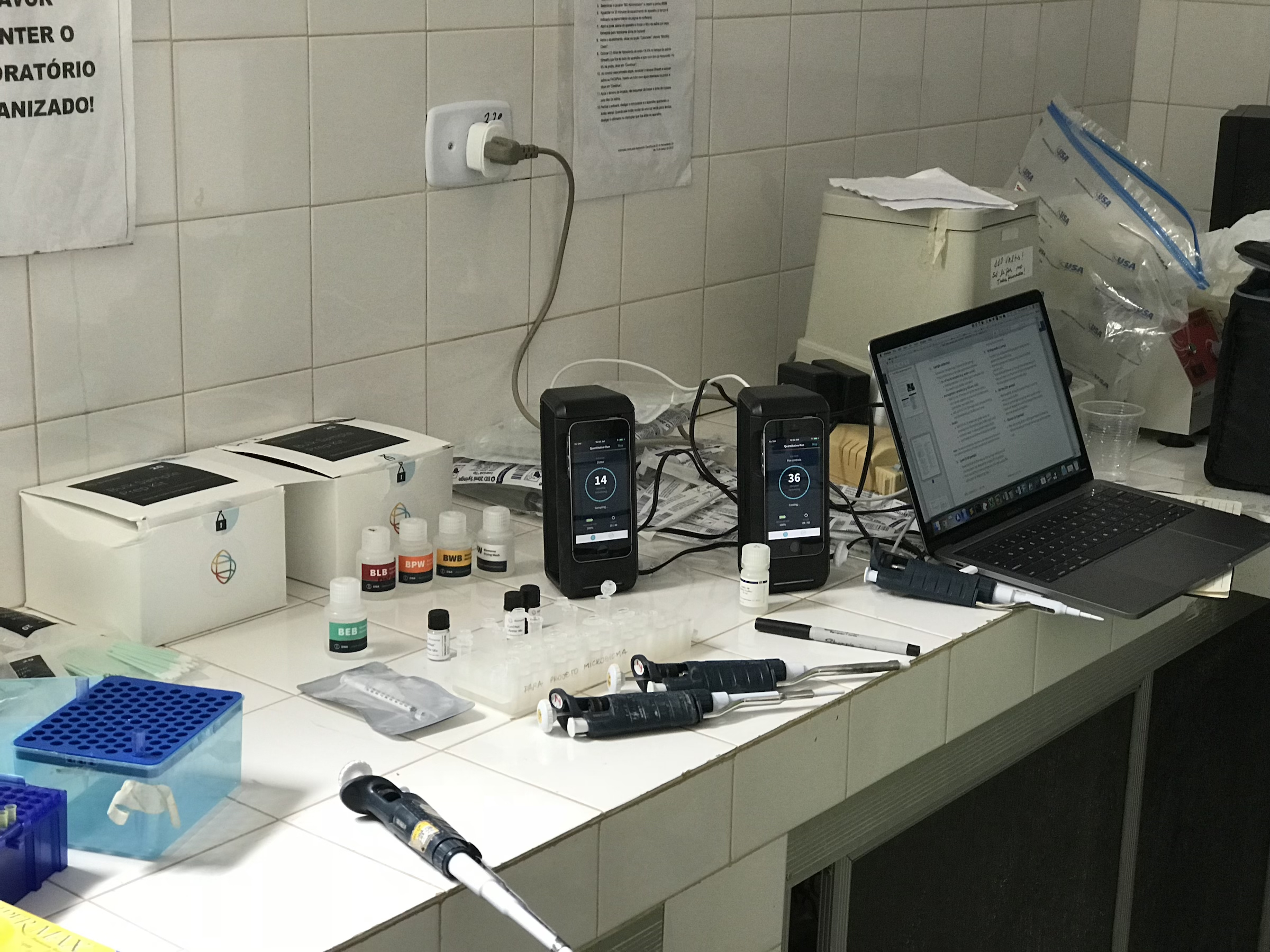
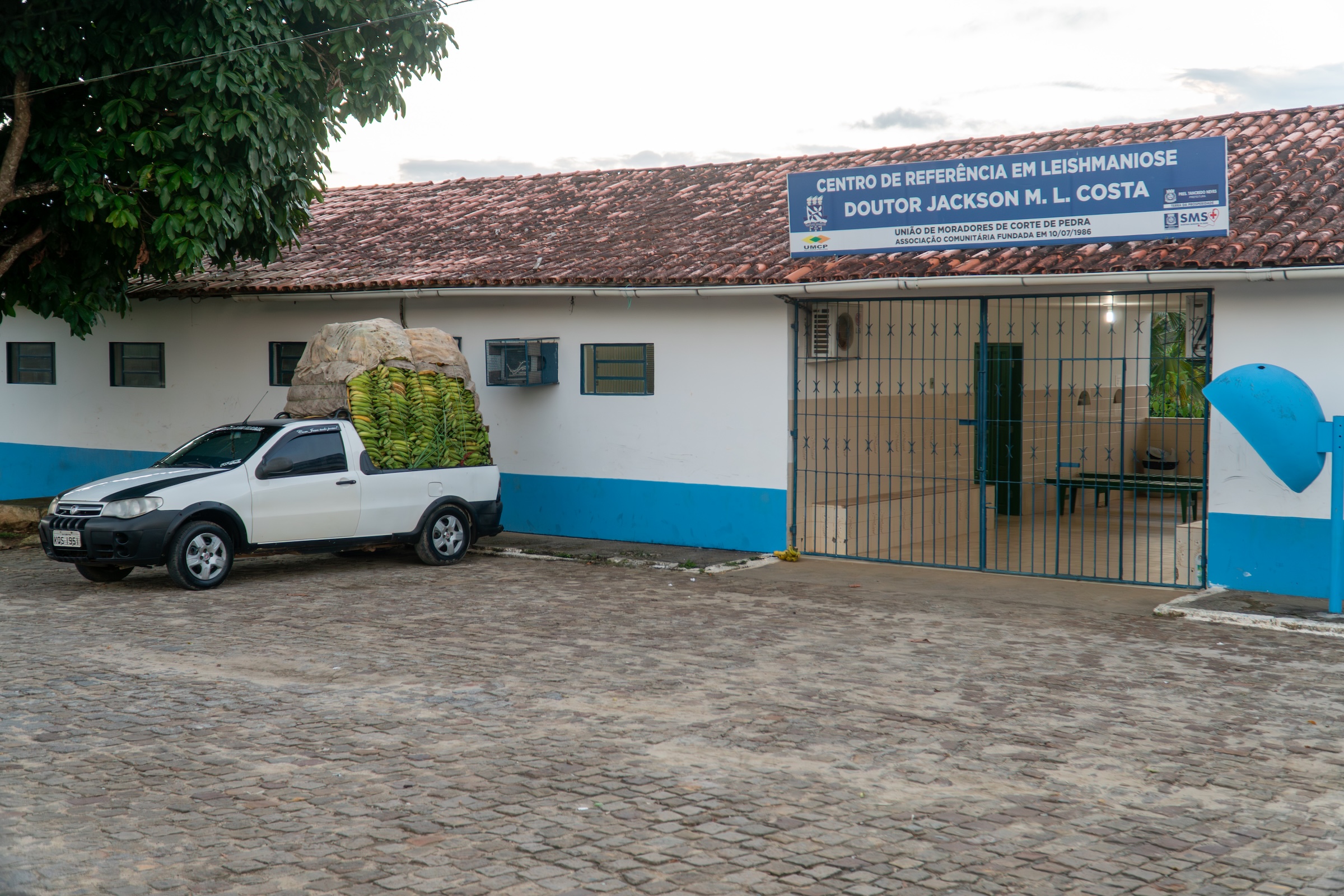
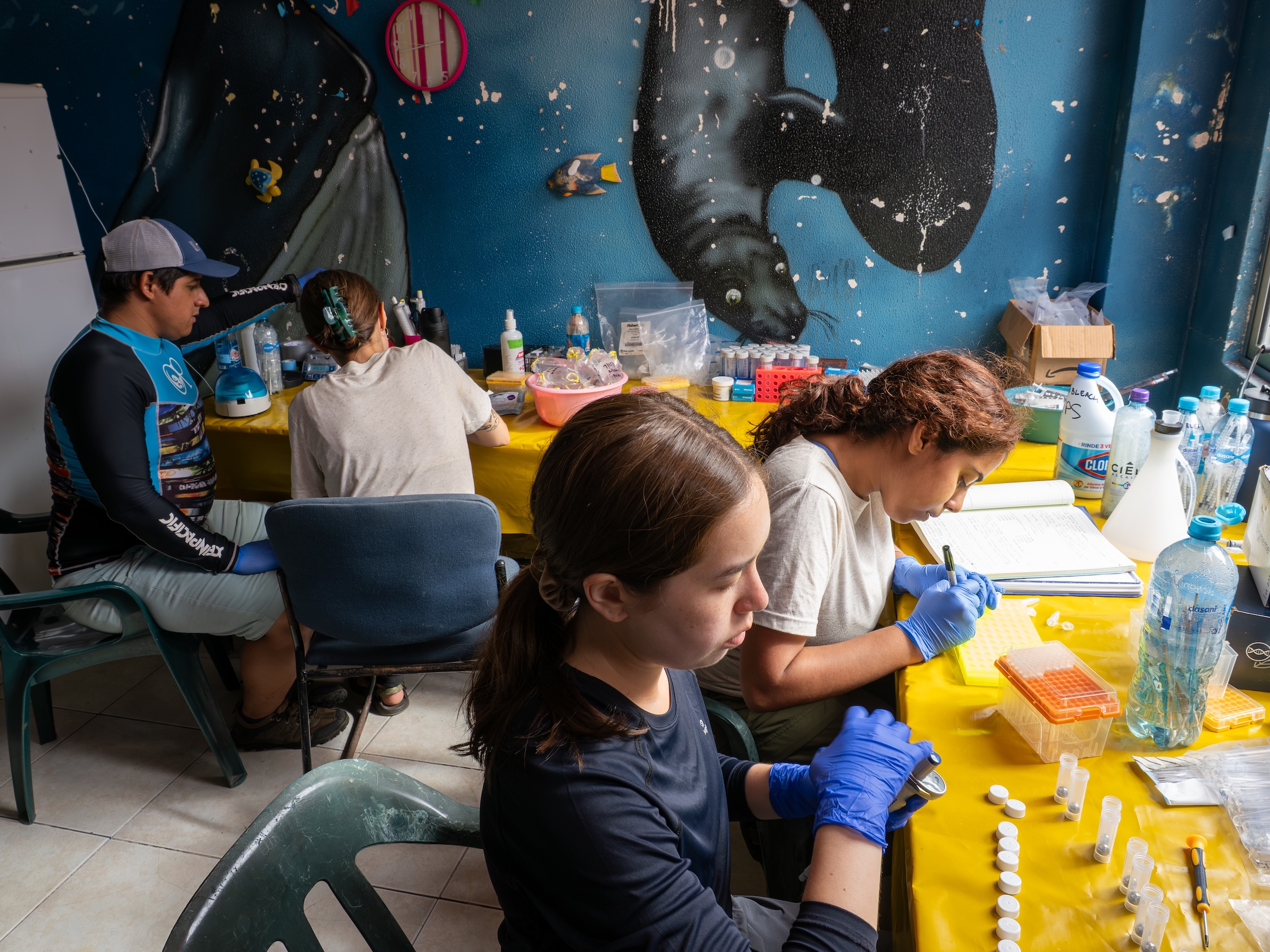
Camila initially came to spend a year in the lab as a PhD student participating in Brazil’s ‘sandwich’ program. During this time, she fell in love with bioinformatics and returned after her PhD to start a postdoc jointly between our lab and the laboratory of Phil Scott. During her postdoc, she took the lead on advancing our understanding of the transcriptional response that drives skin pathology during cutaneous leishmaniasis. Her initial studies identified key biomarkers that predict patient outcomes even before the first treatment has begun, and more recently she demonstrated that localized skin infection with Leishmania elicits a chronic systemic inflammatory response in patients. In her most recent work which involved a collaboration between our lab, the Scott lab, and Elizabeth Grice’s lab, Camila identified a link between skin commensal microbes, in particular Staphylococcus aureus, and slower healing time in cutaneous leishmaniasis. You can read more about that work in this press release. Camila left PennVet in 2023 and is currently a Senior Scientist in Computational Biology and Data Science at Century Therapeutics.
Olivia was a PhD student in the lab and a member of the Microbiology, Virology and Parasitology (MVP) Graduate group from 2019 to 2024. Olivia’s project was focused on the development of genomic methods to study the skin-dwelling parasite, Leishmania braziliensis. In her first paper, she successful developed Selective Whole Genome Amplification (SWGA) for Leishmania species and used this technique to investigate the population biology of this parasite in South America. This work constitutes a breakthrough for the leishmania research field since it allows researchers to amplify whole Leishmania genomes directly from patient material to carry out population genetic studies of this important pathogen, without the need to culture or enrich parasites. Olivia also wrote the first review article on SWGA, which laid out how this method has impacted population genomic studies for complex pathogens.
Pagination