Kat is a VMD-PhD student at PennVet in the Biology Graduate Program. Before coming to Penn, Kat completed her Master’s degree in Evolutionary Anthropology at UC Davis where she studied the antiquity of infectious veterinary diseases in the Peruvian Andes and developed a histological method to identify ancient disease processes in preserved soft tissues. Her current research interests are seated in understanding evolutionary disease ecology and the processes that give rise to modern infectious disease landscapes. Kat is currently rotating in the lab and is working on one of our newest projects, in collaboration with the PA Game Commission, examining bacterial-fungal interactions in the skin microbiome in bats affected by White Nose Syndrome. After grad school, Kat hopes to pursue a residency in wildlife anatomic pathology. When not in class or in the lab, she keeps quite busy putting out fires started by one of her three dogs or three cats.
Rui Xiao is a bioinformatician who joined the lab in June 2024 after earning his Ph.D. in Bioinformatics from the University of Georgia under the mentorship of Dr. Jessica Kissinger. Rui’s PhD work made important contributions to elucidating the transcriptional and epigenetic landscape of the protozan parasite, Cryptosporidium parvum. Since joining the lab, he has been instrumental in developing and optimizing bioinformatics workflows using platforms such as Airflow and Nextflow. Additionally, Rui has a keen interest in studying and fine-tuning local large-language models. Outside the lab, Rui enjoys running and virtual reality fitness training.
Andrew is a bioinformatician that joined the lab in May 2024. He received his PhD in immunology from the University of Pennsylvania where he studied T cells in the contexts of autoimmunity and intestinal homeostasis. In the lab, Andrew specializes in bioinformatic analyses of single-cell and bulk sequencing datasets. He currently works on a large collaborative effort to interrogate intestinal immune responses to varied pathogens at the single cell level. He has an interest in public health and spends his free time reading, baking, and playing volleyball.
Karen is a MD and Pediatric Infectious Disease Society (PIDS) fellow in the Division of Infectious Diseases at the Children’s Hospital of Philadelphia (CHOP). As part of her fellowship, she is spending 3 years in the lab learning about bench research. She has taken the lead on an exciting new project for the lab, which involves testing whether the microbiome may be linked to severe pneumonia in children from lower- and middle-income countries. To address this important question, Karen is using metagenomic profiling of archival samples from the Pneumonia Etiology Research for Child Health (PERCH). Karen is also pursuing a ‘distance-learning’ masters degree in Epidemiology through the London School of Hygiene and Tropical Medicine (LSHTM). These combined efforts underscore her passion for negelected tropical diseases and global health.
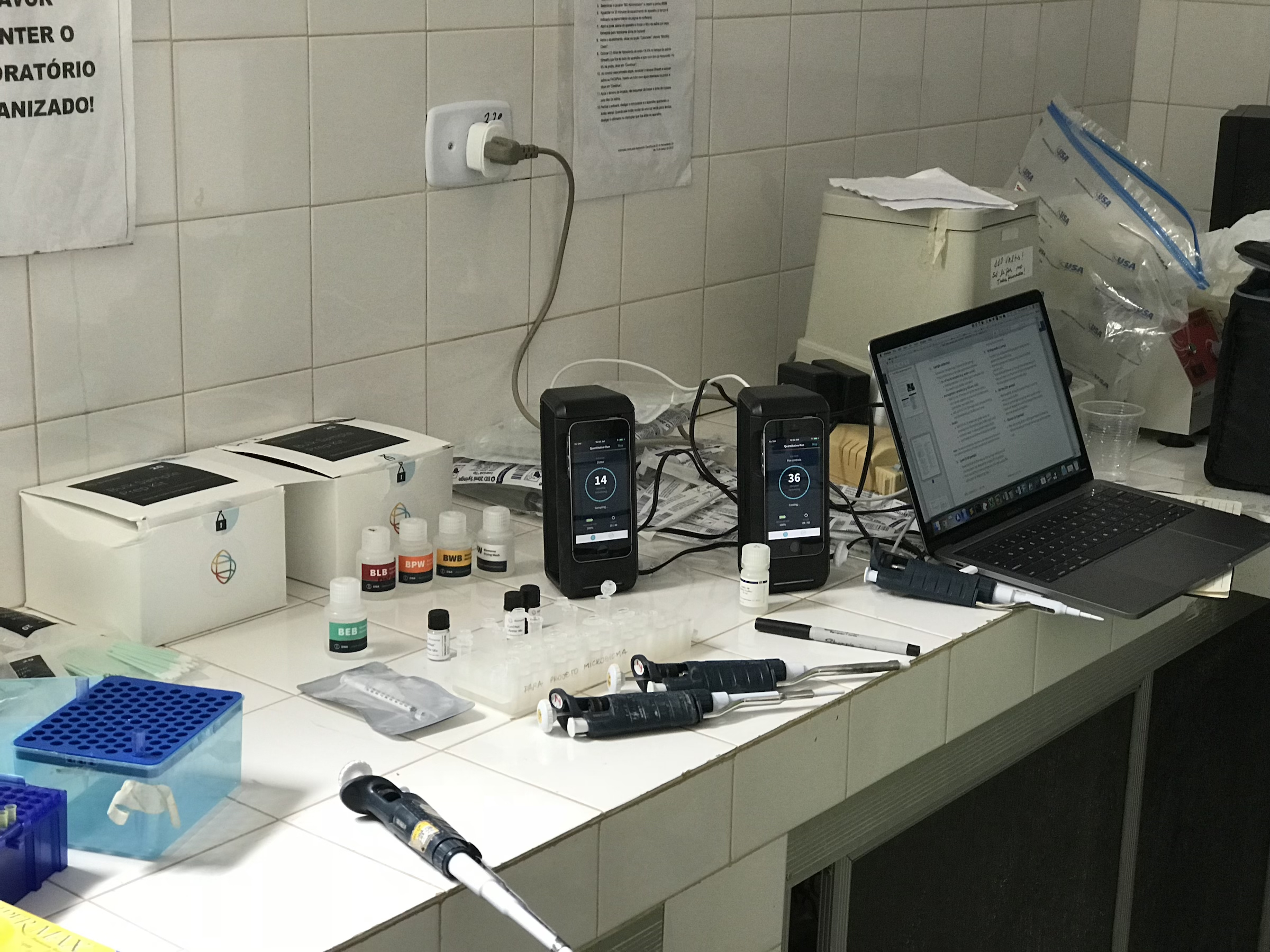
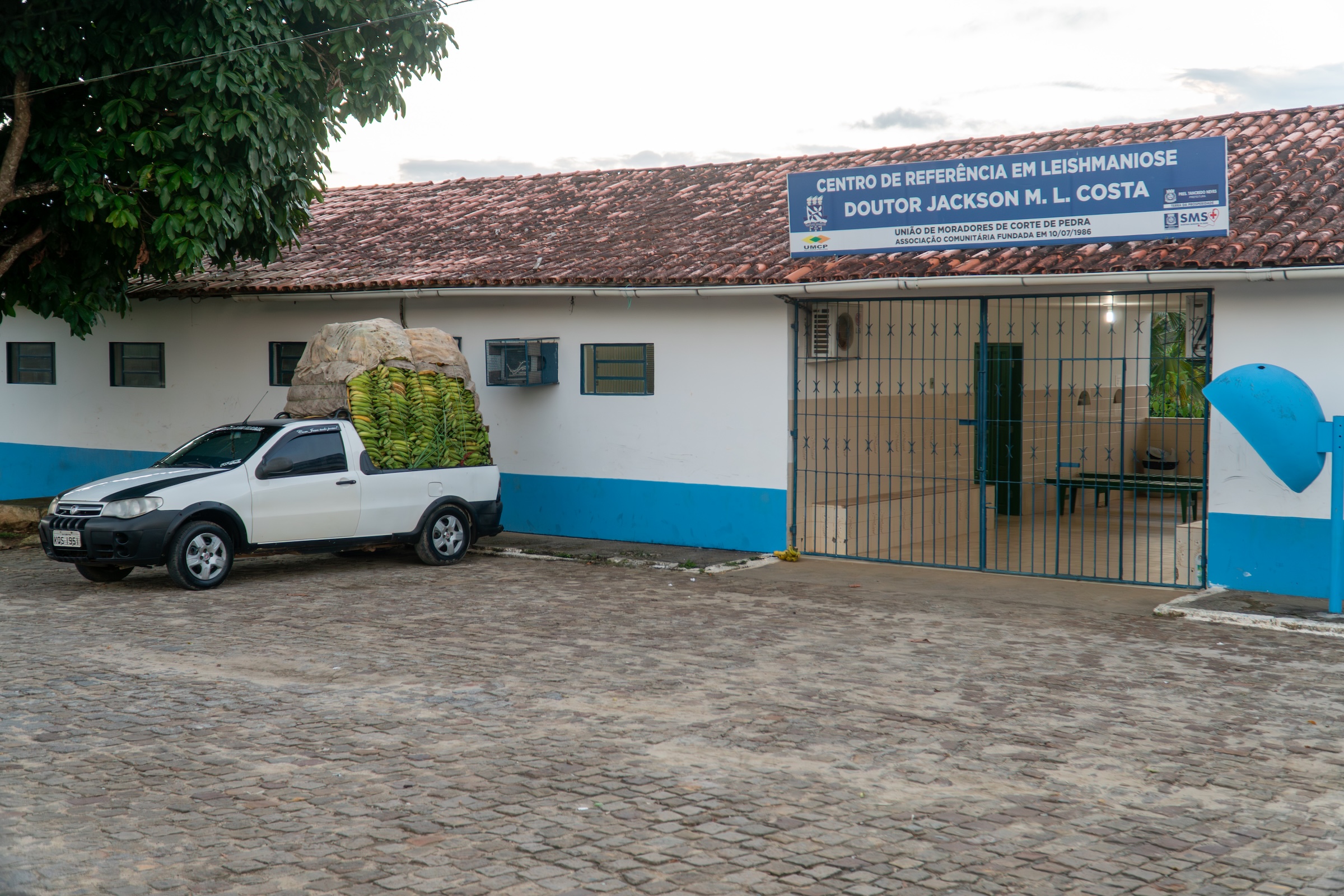
Felipe graduated in Dentistry and has a Master’s degree in Biosciences and Biotechnology from the University of São Paulo, in Brazil, where he’s currently a PhD student. Felipe has been working for over 10 years in the characterization of Salmonella serovars of importance in the “One Health” context. He spent 6-months as a Brazilian “sandwich” student during his PhD. During his time in the lab, Felipe learned computational methods to analyze RNA-seq data with the goal of identifying differences in the gene expression of Salmonella Infantis under conditions of stress. As a side project, Felipe applied his microbiology expertise to our White Nose Syndrome (WNS) project. He was the first person in the lab to begin culturing the causative agent of this disease, the fungus Pseudogymnoascus destructans, and his work laid an important foundation for some reductionist in vitro assays that we are developing to study WNS. In his free time, you’ll probably find Felipe cooking, reading thriller books or watching Marvel movies.
Celine joined the lab in the Summer of 2023 as our first undergraduate through the First Exposure to Research in the Biological Sciences (FERBS). She helped with the early planning stages of a large project the lab began to use metagenomic sequencing and immune profiling of samples from three African sites that participated in the PERCH study.
Samy spent the Fall of 2022 working in the lab to optimize our Selective Whole Genome Amplification work with graduate student, Olivia Pilling. After graduation from Penn, Samy went on to complete a master’s degree in Philosophy of Bioscience Enterprise at the University of Cambridge. Samy was a competitive rower during her time at Penn, and she joined the Australian national rowing team after graduation. She was selected as an alternate for the Australian rowing team for the Paris 2024 Olympic Games.
Yirui (pronounced ‘E-Ray) rotated through the lab in the Fall of 2022 and worked on developing methods for carrying out RNA-seq on the model archeaon, H. volcanii. Yirui is now a graduate student in the lab of Mecky Pohlschroder in the Biology Department at UPenn.
Theresa is a rising second-year VMD-PhD student at Penn and a member of the MVP graduate group. She graduated with a BS in Animal Science from the University of Vermont, where she completed an undergraduate research thesis investigating bovine teat skin commensals with antimicrobial activity against pathogens responsible for mastitis. After college, Theresa spent a year teaching English in southern France before returning to the US to work as a research technician at the University of Wisconsin-Madison. There she worked under the direction of Dr. JP van Pijkeren on several projects revolving around ecology of the gut symbiont Lactobacillus reuteri and the development of probiotic bacteria as delivery vehicles for recombinant biotherapeutics. Broadly, she is interested in small animal internal medicine, the link between diet and microbiome health, and targets for therapeutic manipulation of the microbiome. During her Spring independent study and subsequent Summer rotation, Theresa took the lead on assembling a pangenome for elusive, but important, bile acid-producing commensals. She used the Anvi’o suite of computational tools to carry out comparative analysis of different bile acid-producer species, and to identify bacterial factors that might drive host adaptation.
Shelby is an veterinary student at UPenn (Class of ‘26) who spent the summer of 2023 and 2024 in the lab working jointly with the PennVet Wildlife Futures program. As a ‘WFP’ student, Shelby split her time between bench work and field work. She collected and identified hundreds of ticks recovered from White-tailed Deer (Odocoileus virginianus) in PA state. She extracted DNA from these ticks and ran QPCR tests for a range of pathogens, then used this data to select ticks for metagenomic sequencing.
Pagination